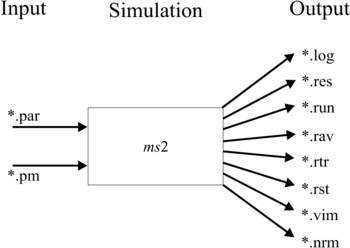
Input and Output Files
The structure of the input files as well as the output files is shown schematically.
Input files
The simulation tool ms2 requires one main input file (*.par) to specify the simulation parameters and at least one molecular force field model file (*.pm) for the considered species.
The *.par file contains all input variables for the simulation such as simulation type, ensemble, number of equilibration and production steps, time step length, and many more. Furthermore, the user can specify temperature, density, and composition.
A *.pm file contains the molecular force field model for a given substance. It contains the parameters and relative position of all sites.
Output files
ms2 yields seven output files:
- *.log file - contains the history of the simulation steps
- *.res file - contains the results of the simulation in an aggregated form. The data is written to the file in reduced quantities as well as in SI units, along with the statistical uncertainties of the calculated properties. The *.res file is created during production and updated a number of time steps or cycles specified in the *.par file.
- *.run file - contains the sampled properties of the simulation for a specified time step or cycle interval. The file is in tabular form, where the data is given in reduced units. The file is subsequently updated according to the user specification, which is specified in the *.par file.
- *.rav file - contains the running averages of sampled properties. The block size and therefore the number of output lines in the *.rav file is user-defined and can be specified in the *.par file.
- *.rst file - is the restart file of the simulation. It contains all molecular positions, velocities, orientations, forces, and block averages for the thermodynamic properties. It is written once at the end of the simulation or immediately after having received a termination signal of the operating system. The restart file allows for the continuation of the simulation, necessary e.g. in case of an early interuption of the simulation, time limits on a queuing system, or unexpected halts.
- *.vim file - contains information on the atomistic configuration, i.e. the positions and orientations of all molecules in an ASCII-form. The configurations are written to the file after a user-specified interval of time steps or cycles. The *.vim file is readable by the visualization tool ms2molecules, which is also part of the simulation package.
- *.nrm file - stores the coordinates of the molecule after principal axis transformation.